New Delhi: The B.1.617.2 lineage of the SARS-CoV-2 virus — which was recently identified in India — is now growing in prevalence across the country, genome sequencing data shows.
As of Tuesday, as many as 12,179 genome sequences of the SARS-CoV-2 virus have been uploaded on the GISAID website, a global repository of genomic data.
The SARS-CoV-2 lineage B.1.617 has been dominating the second wave of the Covid-19 pandemic in India over the past few months. B.1.117 — a lineage that was first identified in the UK — also dominated the spread of infections for a while in India. However, B.1.617 soon took the lead, accounting for 29 per cent of the samples sequenced.
Additional sequences uploaded from India show that in the last 60 days, B.1.617.2 has been gaining prevalence in India.
What is B.1.617.2?
While the World Health Organization (WHO) has classified B.1.617 variant of the coronavirus as a ‘variant of concern‘, it appears that this lineage consists of at least 3 sub-lineages — B.1.617.1, B.1.617.2 and B.1.617.3.
The lineage B.1.617 was defined by the mutations E484Q, L452R and P681R -—along with D614G — in the spike protein.
B.1.617.1 has an additional mutation Q1071H in the spike protein. B.1.617.2 is defined by more mutations in the spike protein, namely T19R, DEL157/158, T478K and D950N.
B.1.617.3, meanwhile, has the following additional mutations: T19R, DEL157/158 and D950N.
It is important to note that each of these viral lineages have more mutations, but the ones mentioned above are considered to be more important since they occur in the spike protein of the virus which facilitates the entry of the virus into the host cells.
B.1.617.2 in India
The first sample of B.1.617.2 in India was detected on 12 December 2020. Since then, more than 350 sequences of this lineage have been detected with prevalence increasing to 75 per cent at the end of April.

Last week, the Public Health England named B.1.617.2 a variant of concern as evidence showed that the virus had increased transmissibility.
There is still not enough data to confirm if any of the variants recently detected in India cause more severe disease or render the vaccines currently being administered any less effective.
Meanwhile, the prevalence of B.1.117 has now fallen to below 5 per cent in India. B.1.351 — the variant first detected in South Africa — also has very low prevalence (between 0 to 4 per cent in the month of April).
B.1.617.1 dominates the infection spread, although B.1.617.2 seems to be fast catching up since the last two months.
The current data shows that 52 per cent of the viral genomes sequenced in the last 60 days belonged to the B.1.617.1 lineage, while 20 per cent belonged to the B.1.617.2 lineage.
Maharashtra

Less than five per cent of the samples sequenced in the last two months belong to the B.1.117 lineage in Maharashtra.
B.1.617.3 has a prevalence of 3 per cent in the state.
Prevalence in India over last 60 days
The cumulative data from the last 60 days shows that more than half the sequences recently sampled in India belong to the B.1.617 group.
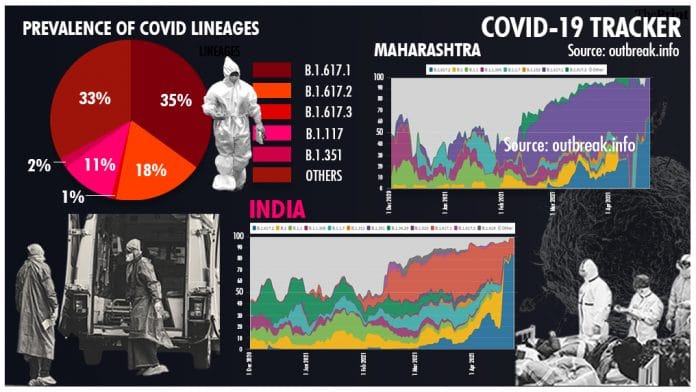
As much as 35 per cent of the total samples are B.1.617.1, 18 per cent are B.1.617.2. B.1.617.3 currently accounts for just 1 per cent of the samples sequenced in the last 60 days.

The data available about the variants in India is very limited, as samples of less than 1 per cent of those infected have been sequenced so far. The understanding of the prevalence of different variants in India is likely to continue to evolve as more and more samples are sequenced.
Also read: Sputnik V arrived 1 May to boost India’s vaccine drive, but is still ‘stuck’ in lab for tests