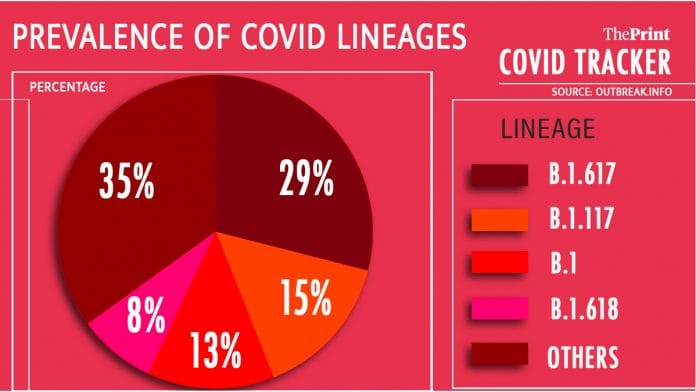
New Delhi: In the last 60 days, the B.1.617 lineage of the SARS-CoV-2 virus has sped ahead in the evolutionary arms race of the viral spread in India, leaving behind almost all other lineages circulating in India.
A lineage is defined by a set of genetic mutations that alter some of the amino acids (the building blocks of proteins) of the original virus.
As much as 29 per cent of the samples sequenced were of the B.1.617, the lineage defined by the mutations E484K, L452R and P681H.
A close second is now B.1.117 — the lineage first identified from the UK — with 15 per cent of the samples sequenced having the mutations characteristic of this lineage.
B.1 — a lineage that was reported early in the pandemic defined by the mutation D618G among others — dominated 13 per cent of the total samples over the last 60 days.
A newly identified lineage — named B.1.618 — seems to now grow in number. In the last 60 days, the lineage was found in 8 per cent of the samples sequenced. ThePrint had reported that most of the samples of B.1.618 had been identified from West Bengal.
The B.1.168 variant is characterised by the deletion of two amino acids in the spike protein, described as H146del and Y145del. Along with this, the lineage also has the mutations E484K and D618G.
Currently, the Indian SARS-CoV-2 Genomic Consortia (INSACOG) has fully sequenced 15,135 samples of the virus. This included sequencing of samples from 1,932 international travellers. Of these, 9,115 sequences have been uploaded on the GISAID website, a global repository of genomic data.
Dr Sujeet Kumar Singh, director of National Centre for Disease Control (NCDC), told ThePrint during a virtual media briefing held Friday that the samples for sequencing are collected from a representative population in any area that is experiencing a surge in cases.
“There are too many cases detected nationally and we simply do not have the infrastructure to sequence each and every sample,” Singh added.
ThePrint looked at the open access data currently available on the genome sequences to ascertain how the prevalence of genome sequences has changed over the course of the pandemic.
However, it should be noted that since India has sampled less than 0.01 per cent of the total number of people infected, there is a possibility that the available data does not give an accurate representation of prevalence of mutations.
Moreover, since the number of samples sequenced has changed over time, the data looks at the proportion of the samples of each lineage, rather than looking at the absolute numbers.
Here, lineages are now defined by the mutations that are of interest, usually the ones that cause changes in the spike protein. However, every lineage likely has more than one characterising mutation.
India
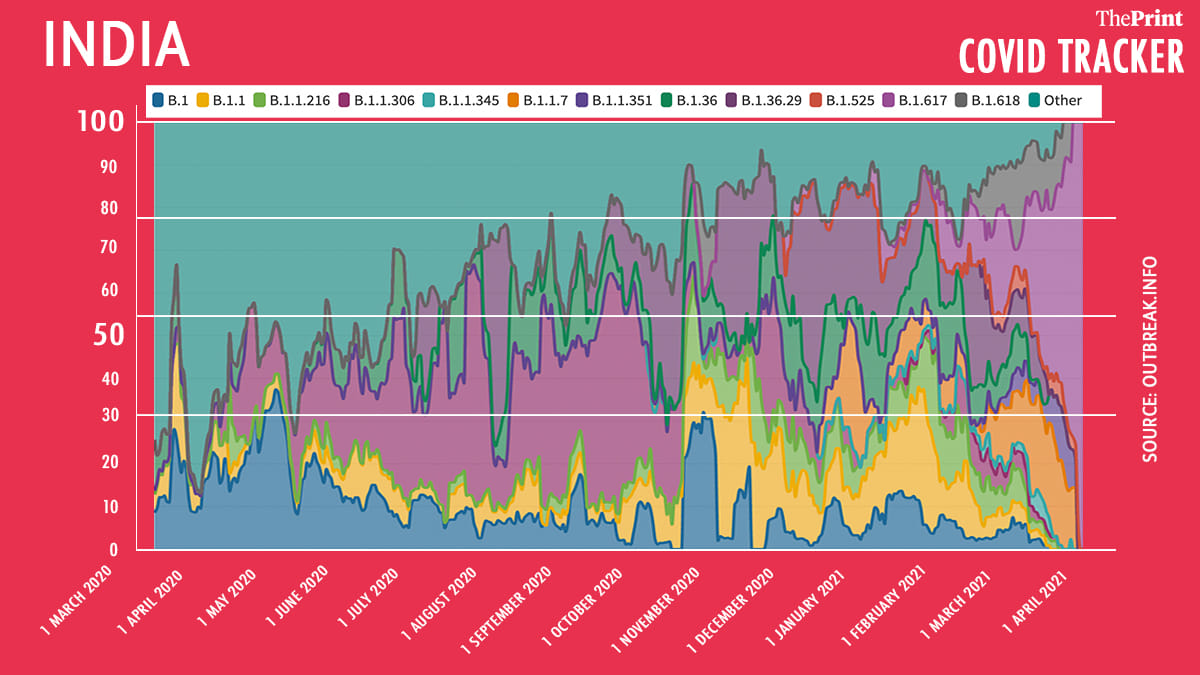
Overall, since the beginning of the pandemic, the mutation that dominated the Indian population was D618G, a mutation in the spike protein that is linked to increased transmission. The lineage carrying this mutation, called B.1, was the most prevalent in the Indian population till about June 2020.
However, in July the lineage designated B.1.1.306 took over. At the peak of the pandemic in September last year, as much as 54 per cent of the samples in India was from this lineage. The mutation of concern in this lineage is also D618G. However, other defining mutations were not deemed of importance for this lineage.
The lineage prevalence fell as the first wave of Covid subsided in India.
Although there was much concern over the B.1.117, a lineage identified in the UK that was found to be more transmissible, its prevalence remained relatively low. This may have been because concern around the mutation kicked into action strict tracing and containment measures, some of which may have helped stop the lineage from spreading further. However, Singh today reported that the spike in Punjab and Delhi can be attributed to this lineage.
B.1.617, the lineage first identified in India, is rapidly growing in prevalence. From about 11 per cent in January this year, the prevalence of this lineage has gone up to over 50 per cent in March.
Due to the lack of adequate data available, it is currently not possible to accurately assess the prevalence of different sequences in different states. However, ThePrint looked at sequences from Maharashtra and West Bengal, the two states from where the maximum number of sequences have been uploaded.
Maharashtra
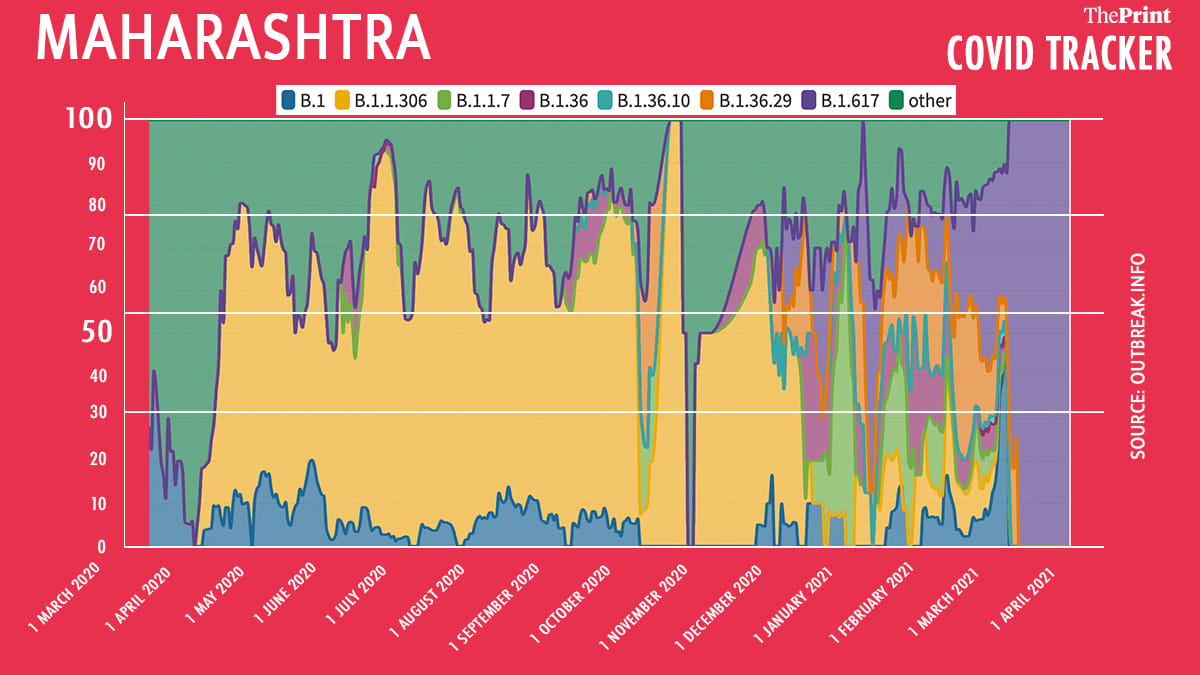
Over 2,000 sequences from Maharashtra have been uploaded on the GISAID platform. Of these, 42 per cent belong to the B.1.617 lineage. The prevalence of this lineage increased from 8 per cent in January to nearly 100 per cent in April. However, this needs to be interpreted with caution since very few samples have been sequenced in the last 60 days.
West Bengal
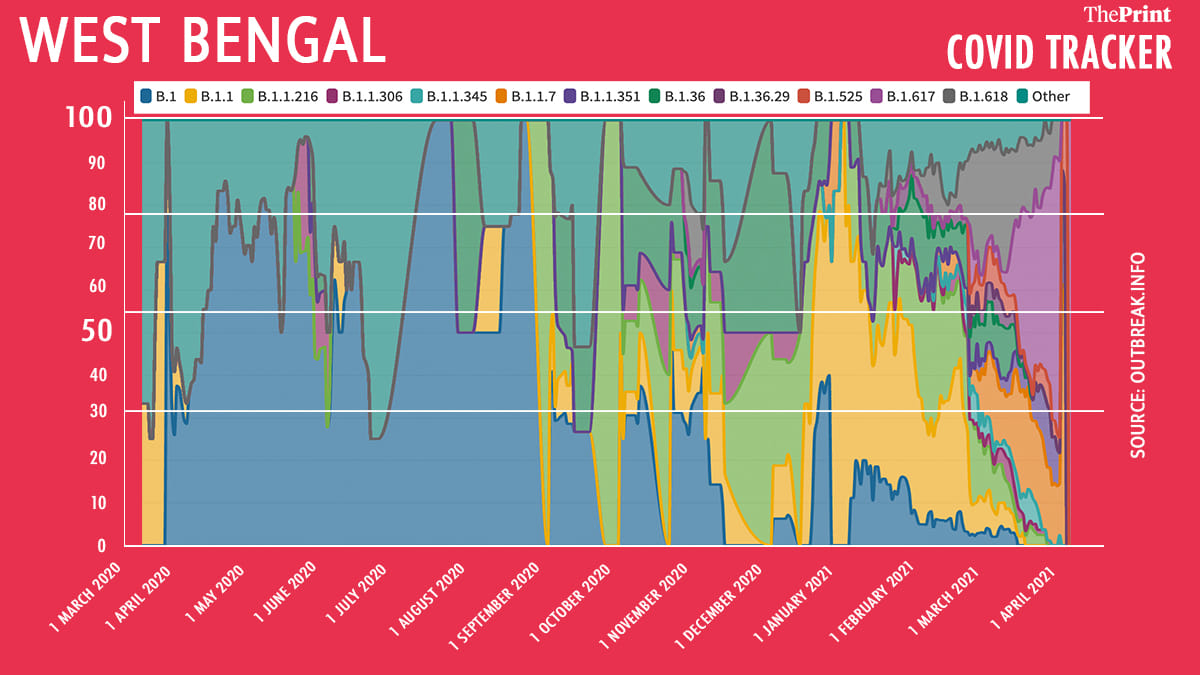
Data of over 1,300 samples from WB have been uploaded on GISAID. Here too, B.1.617 lineage showed up with 28 per cent prevalence. However, B.1.618 lineage has had an 18 per cent prevalence in the state in the last 60 days.
(Edited by Manasa Mohan)